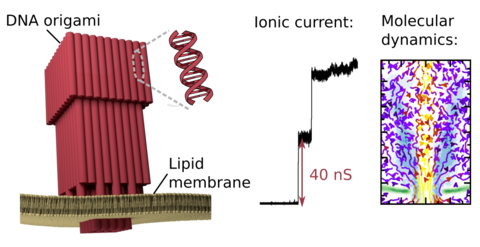
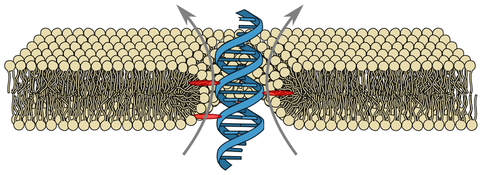
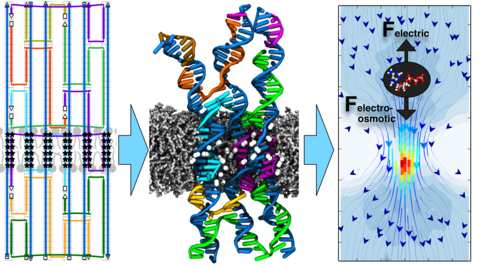
DNA membrane channels
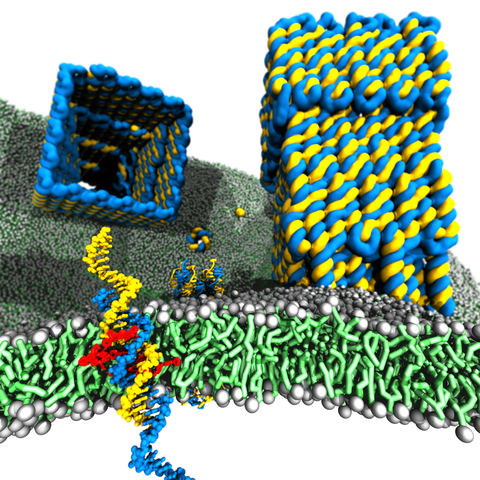
Membrane protein channels involved in cellular signal transductions are fascinating biological sensors with high selectivity and efficiency. Recently, DNA origami nanostructures emerged as highly customizable mimetics of biological membrane channels. A typical DNA membrane channel is assembled from several DNA double helices arranged into a polygon pattern, with the central cavity forming the transmembrane pore. To facilitate insertion of the DNA channel into a lipid bilayer membrane, the DNA helices are chemically modified to carry hydrophobic anchors. Until now, most of the DNA channels featured four or six DNA helices arranged in a square or a hexagon, with the inner channel diameter varying between 1 and 2.5 nm. Using all-atom Molecular Dynamics (MD) simulation, we characterized the biophysical properties of the DNA membrane channels with atomic precision. We further engineered DNA membrane channels that span one order of magnitude in diameter and three orders of magnitude in conductance and molecular weight, covering the entire range of the protein membrane channels.