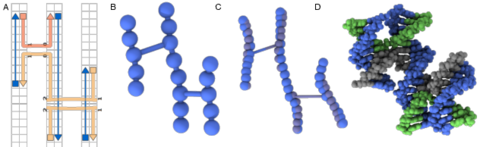Multi-resolution simulations of self-assembled DNA nanostructures
The DNA origami method has brought nanometer-precision fabrication to molecular biology labs, offering myriads of potential applications in the fields of synthetic biology, biomolecular medicine, molecular computation, etc. Advancing the method further requires controlling self-assembly down to the atomic scale. Hence, fast, accurate and detailed structure prediction can advance the development of complex, functional DNA origami constructs. Towards this goal, we developed a protocol that unites the rapid and robust structural relaxation of low-resolution coarse-grained modeling with the detail and accuracy of atomistic molecular dynamics simulation. In this tutorial we will show you how to use the multi-resolution DNA (mrdna) Python package to perform such simulations.