Introduction to MD simulations of Plasmonic Nanotweezers
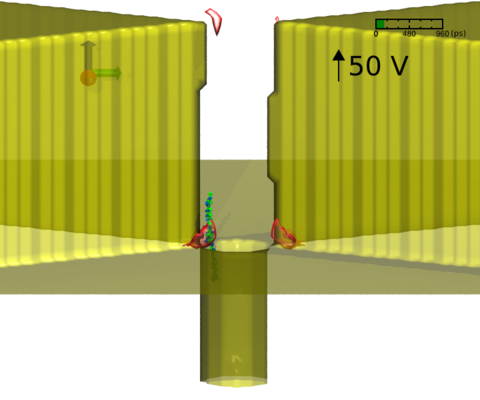
In this tutorial, we will demonstrate how to manipulate the translocation of a polypeptide through a solid-state nanopore with a plasmonic field in MD simulation. We will start from obtaining the 3D plasmonic field distribution using nanoDDSCAT+ (https://nanohub.org/tools/ddaplus) on nanoHUB. Then, apply the 3D plasmonic field distribution in MD simulation to trap the translocation of a charged polypeptide in an electric field using NAMD (http://www.ks.uiuc.edu/Research/namd/), a parallel molecular dynamics simulation program. Finally, we will use a molecular visualization software VMD (http://www.ks.uiuc.edu/Research/vmd/) to see the simulation trajectory and analyze the result. It is assumed that the reader of this tutorial has some basic knowledge about interacting with the UNIX command-line interface, like cd, ls, mkdir, cp and so on.