A tetrahedral DNA nanorobot with conformational change in response to molecular trigger
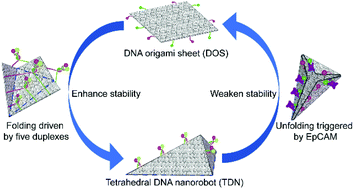
Dynamic DNA origami nanostructures that respond to external stimuli are promising platforms for cargo delivery and nanoscale sensing. However, the low stability of such nanostructures under physiological conditions presents a major obstacle for their use in biomedical applications. This article describes a stable tetrahedral DNA nanorobot (TDN) programmed to undergo a controlled conformational change in response to epithelial cell adhesion molecule (EpCAM), a molecular biomarker specifically expressed on the circulating tumor cells. Multiresolution molecular dynamics simulations verified the overall stability of the folded TDN design and characterized local distortions in the folded structure. Atomic force microscopy and gel electrophoresis results showed that tetragonal structures are more stable than unfolded DNA origami sheets. Live cell experiments demonstrated the low cytotoxicity and target specificity of TDN. In summary, the proposed TDN can not only effectively resist nuclease catalysis but also has the potential to monitor EpCAM-positive cells precisely.